Better Medicines thanks to IT
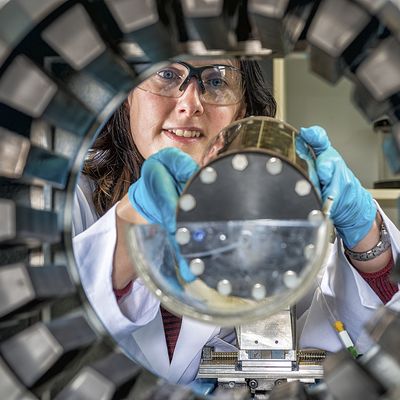
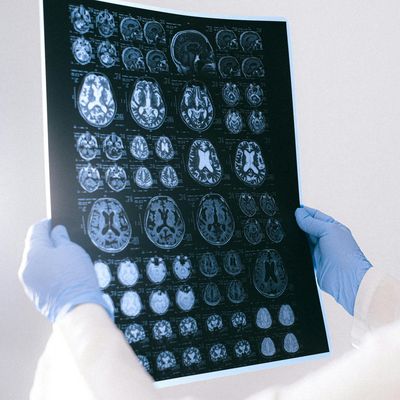
For the development of new medical agents it is important to know how biomolecules, for example viruses, are structured. Bioinformatician Anna Theresa Cavasin therefore wants to use computer-assisted methods to optimize images from cryoelectron microscopy - in order to pave the way for better drugs with data science.
The bicycle racks in front of the Centre for Structural Systems Biology, or CSSB for short, stand empty on this sunny fall morning. Offices and labs at the interdisciplinary infection research center in the Bahrenfeld quarter of Hamburg are sparsely occupied due to the coronavirus pandemic. Anna Theresa Cavasin, a bioinformatician and doctoral candidate, is wearing a face mask when she comes down to the building’s revolving door; she then leads us up an elegantly curved staircase made of light wood to the conference area on the second floor. We meet a few colleagues on their way to the coffee machine, but otherwise, everything is quiet in the bright new building that opened in 2017 here on the German Electron Synchrotron (DESY) campus.
The CSSB actually has enough space for 180 scientific staff. In more normal times, Anna Theresa Cavasin works three days a week here together with other researchers in one of the open-plan offices connected to the lab. The 26-year-old is wearing black jeans and a checkered flannel shirt; her gaze is focused and alert. For nine months now, Cavasin has been one of 14 doctoral candidates at the DASHH graduate school, which is officially known as Data Science in Hamburg – Helmholtz Graduate School for the Structure of Matter. The school—part of the Helmholtz Information & Data Science Academy (HIDA), Germany’s largest postgraduate training network in the field of information and data science—is a hub for projects led by young scientists who conduct basic research into the structure of matter. Cavasin’s project is called “Next generation integrative modeling for cryoelectron microscopy.”
The goal: to improve the microscopic image quality of biomolecules
While its title might sound somewhat abstract, the project pursues a very tangible goal: Cavasin wants to improve imaging results in cryoelectron microscopy, an increasingly relevant method of determining the structure of biomolecules. In simple and specific terms, she does this by applying an algorithm to automatically sort out poor-quality results with little informative value from the vast quantities of images that are produced when working with a microscope. In the over nine months since she became a doctoral candidate at DASHH, the young bioinformatician has been working to select parameters that differentiate good images from those of poor quality. Once she has determined these variables, Cavasin will spend the remaining part of the three years planned for her doctorate working to program algorithms that sort image material accordingly.
This process is somewhat more complex in scientific terms, of course. So, it’s worth taking a look at the starting point of Cavasin’s research. Understanding the structure of certain biomolecules to the most accurate extent possible is a basic area of knowledge that is required in the development of drugs in particular. Cryoelectron microscopy is increasingly being used to arrive at valid data relating to these smallest of particles. Cavasin works with frozen rather than crystallized samples—which, among other things, has the advantage of enabling the analysis of proteins that cannot be crystallized. The resolution of the images produced by cryoelectron microscopes has increased significantly over the past ten years—in fact, those responsible for developing the method were awarded the Nobel Prize for Chemistry for their contributions in 2017. But, Cavasin says, there are still many cases in which the quality of the images is insufficient.
Why would scientists need hundreds of thousands to millions of 2D microscopic images of a single molecule? Because they are the basis for the incredibly large-scale process of calculating 3D models—for instance, of a certain protein. One thing is for certain: The better the individual 2D images, the more exact the 3D image produced from them. And what happens next with the 3D model? Cavasin explains: “We want to see what a protein involved in an infection looks like, down to the level of individual atoms if possible. This is the only way we can develop active ingredient molecules—in other words, drugs—against a virus, for example.”
What really motivated her, were her friends
Even when she was in school, Cavasin was very interested in biology and chemistry, and especially in chemical and molecular biology. She entered school competitions together with two of her friends, who are now working toward their doctorates in the sciences as well. On the website of her school in Essen, where she grew up, there are still images of her smiling into the camera as she holds a certificate. She carried out various analyses back then, including for her presentation at the International Biology Olympiad, “Onion Cells in the Ion Trap”—which earned her a place among the top twelve participants from North Rhine-Westphalia. Did her teachers notice and encourage her talent? Not really, Cavasin says; she did a lot of reading herself, in books and online. But what really motivated her, she says, were her friends. They were also the ones who drew her attention to the opportunity of starting university early when she was 17. And this was how Cavasin started regularly attending university lectures in her last year of high school rather than English classes, which she describes as boring.
"I think informatics is the best way to approach questions in biology."
Anna Theresa Cavasin, doctoral researcher
After completing high school, she then began studying toward a bachelor’s degree in chemical biology in Dortmund. Given where she is today, it’s clear that her thesis on computer-assisted active ingredient design determined her next move, which took her to Hamburg for a master’s degree in bioinformatics. Cavasin studied under Matthias Rarey, Head of the Center for Bioinformatics at the University of Hamburg—and one of the spokespersons for the new DASHH graduate school. Rarey is now supervising Cavasin’s doctoral thesis as well. In addition to her desk at CSSB on the DESY Campus, she also has a workstation at the university. She says that working in bioinformatics lets her get to the heart of what she’s really interested in: “I think informatics is the best way to approach questions in biology, because it’s so structured. I like structures. And I like efficiency.” A sense of curiosity is something that her group at the graduate school shares as well, she says, noting that their main question is always: What’s the reason for that? “Gaining new knowledge is our common goal.”
Cavasin says that she sees the world in a rational way. She wants to understand—down to the smallest detail, wherever possible. “I’ve always been interested in what’s behind processes. Why the world works the way it does.” She’s happy that she has ended up doing her doctorate in the special field of cryoelectron microscopy. “The job we do here isn’t one that a lot of people do—and I think that understanding something like this at a very fundamental level at some point is amazing. But only as long as I feel that what I’m doing is worthwhile—and that my work ultimately helps people in a tangible way.” Such as giving them access to better drugs, for example.
Author: Christiane Langrock-Kögel